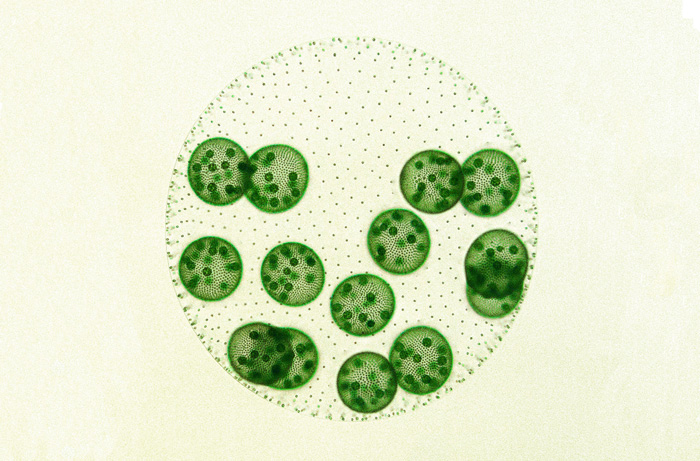
Recent evolution of simple germ–soma division in a green alga sheds light on the early stages of complex multicellular life.
Among evolution’s greatest innovations are germ cells. These specialized reproductive cells—familiar to us as sperm and eggs in humans—set the stage for complex multicellular life because they free up all the other cells in the body (known as somatic cells) to specialize for many other functions. Because they appeared so long ago in our evolutionary history, the way our germ cells emerged has been obscured, leaving many questions about this momentous biological turning point.
We may never know precisely how the germ–soma dichotomy arose in our lineage, but scientists are still searching for clues. In a report in G3, researchers Gavriel Matt and James Umen describe their work on the green alga Volvox carteri, an intriguing species that has recently and independently evolved a simple germ–soma division. In this alga, a spheroid composed of somatic cells and extracellular matrix surrounds germ cells called gonidia. The gonidial cells undergo embryogenesis to produce new juvenile spheroids that hatch from their mother spheroid, whereas the somatic cells that are left behind eventually senesce and die. This system offers a unique opportunity to explore the early evolution of germ cells.
By comparing the V. carteri germ and somatic cell transcriptomes, Matt and Umen found that the somatic cells had more transcripts from young, lineage-specific genes, whereas the germ cells had more transcripts from ancient genes that are similar to those expressed in stem cells from animals and land plants. The germ cells also expressed more genes overall than the somatic cells did, despite being specialized for reproduction. Although counterintuitive, this is reminiscent of the way certain pluripotent stem cells in other systems express a greater number of genes—including more ancient genes—than their somatic daughter cells do.
The duo also investigated the idea that V. carteri evolved its germ–soma division by repurposing genes initially used for the transition to a simpler form of multicellularity without a germ–soma dichotomy, but found little support for this hypothesis. Genes exclusive to V. carteri and one of its close multicellular algal relatives that does not have differentiated cell types, Gonium pectorale, were not specifically enriched in either V. carteri cell type. Instead, they discovered that each V. carteri cell type expresses orthologs of genes from different temporal phases of the light–dark cycle in a related unicellular alga, Chlamydomonas reinhardtii, with somatic cells enriched for expression of dark-phase genes, and gonidial cells enriched for expression of light-phase genes. These results suggest that V. carteri cell types may have evolved through cooption of temporal gene regulation in an ancestor whose different phases were converted into germ- and soma-specific expression programs.
Notably, the researchers also found increased expression of genes involved in anabolic pathways in gonidial cells, which contrasts with the upregulation of genes involved in catabolic pathways found in somatic cells. Specifically, somatic cells preferentially expressed genes involved in breaking down carbon stores into sugars, which are important building blocks for the extracellular matrix that surrounds gonidia and provides a structural scaffold that maintains organismal shape and integrity. These observations suggest that the somatic cells sacrifice themselves for the multicellular collective, contributing stored resources at their own expense. With such an investment in the next generation, it’s no surprise that germ–soma dichotomies have evolved repeatedly, giving researchers fertile ground to explore the rise of complex lifeforms.
CITATION:
Cell-Type Transcriptomes of the Multicellular Green Alga Volvox carteri Yield Insights into the Evolutionary Origins of Germ and Somatic Differentiation Programs
Gavriel Y. Matt and James G. Umen
G3: GENES|GENOMES|GENETICS 2018 8: 531-550; https://doi.org/10.1534/g3.117.300253
http://www.g3journal.org/content/8/2/531