In this month’s editorial, the Editors of GENETICS invite submissions of human genetics research articles. To kick off the journal’s call for papers, the October issue features an article by Brooks and Wall et al. identifying the cause of a single-family disorder and a commentary by Phil Hieter and Kim Boycott on the power of model organisms for understanding rare disease pathogenesis. If you’re at the 2014 ASHG meeting, stop by the GSA booth and pick up a copy!
Susan Brooks had reached a dead end.
Brooks, a medical geneticist at Rutgers Robert Wood Johnson Medical School, was trying to diagnose a boy who had a genetic disorder she had never encountered before. The young patient showed delayed development, microcephaly, recurrent febrile illnesses, seizures, overall slow growth, and several minor craniofacial and limb malformations.
She strongly suspected an X-linked mutation, after finding that two of the boy’s maternal uncles had similar unexplained neurodevelopmental disorders. In addition, the boy’s mother and maternal grandmother displayed non-random patterns of X-inactivation, a clear sign that one of their X-chromosomes carried a harmful mutation.
But the results of a clinical sequencing panel for X-linked intellectual disorders did not fill Brooks with confidence. Of the 81 genes in the panel, the only mutation carried by the boy that was not present in healthy populations was in RPL10, which encodes a component of the large ribosomal subunit. RPL10 was included in the panel because other variants had been implicated in two cases of familial autism, but those cases were nothing like this family’s disorder. Mutations in other ribosomal protein genes can cause Diamond-Blackfan anemia, but this family showed no signs of being anemic.
It was not even clear whether the new variant—a missense mutation—was deleterious to Rpl10 function, as the bioinformatic predictions were ambiguous.
All in all, Brooks needed more proof that the RPL10 mutation was causative. Unless someone stumbled across another family with the same symptoms, this tantalising clue had turned into a dead end.
Then, in 2012, she attended a meeting of the Human Genetics Association of New Jersey, where the invited speaker was Erica Davis from the Center for Human Disease Modeling at Duke University.
The bottleneck in disease genetics, Davis told the audience, is functional annotation. Clinical sequence is accumulating at a rapid rate, but often provides only ‘variants of unknown significance.’ For sequence information to be useful to patients, clinicians need systems for efficiently testing relevance of gene function to disease phenotypes and analyzing the pathogenic potential of mutations. That’s where model organisms come in, said Davis. She
and her colleagues specialize in using zebrafish to provide crucial functional information on variants associated with rare diseases, even those that occur in just one pedigree. One-of-a-kind families, she calls them.
Brooks approached Davis after the talk with the story of the family with the ambiguous mutation. Davis agreed to help. It was just the right task for her new sophomore student, Alissa Wall, who would need a well-defined project for learning the ropes of zebrafish work.
Remarkably, when Wall knocked down the zebrafish RPL10 ortholog, the fish had normal body length but significantly smaller head size. Expression of human RPL10 rescued the phenotype, but human RPL10 bearing the clinical mutation did not. In short, the RPL10 variant caused the zebrafish to develop microcephaly. Not only did Wall’s research generate data for an award-winning honors thesis, it also provided good evidence that it is a loss-of-function mutation and is likely to be responsible for the family’s genetic disorder.
Important as this evidence is, it’s far from a “cure” and has no obvious implications for medical management of the disorder. But it’s a start, and the group has already made progress in understanding how loss of Rpl10 function causes small head size in fish. The lab across the hall from Davis (led by
Christopher Nicchitta) studies mRNA translation, and they helped show that RPL10 knockdown decreases bulk translation in the head and increases apoptosis in the brain. Davis and Nicchitta are pursuing these leads further, hoping to turn up something clinically helpful. For example, they are interested in whether the growth phenotypes are caused by the overall decrease in translation in the brain, or whether the critical trigger is deficiency of a particular subset of proteins.
This research also revealed a new class of ribosomopathy. Now, other components of the large ribosome subunit may be added to the list of candidate genes for rare causes of microcephaly. Thanks to a diverse partnership — patient families, clinical geneticists, zebrafish researchers, and biochemists — Brooks’ dead end is turning into a new beginning.
Read this month’s GENETICS editorial calling for human genetics article submissions.
Read Phil Hieter and Kim Boycott’s commentary on model organisms in rare disease genetic
research.
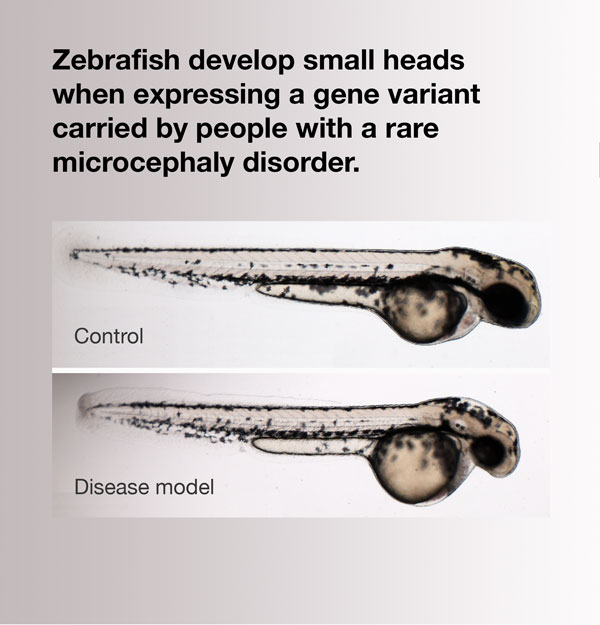
When the gene mutated in patients with a rare disorder is knocked down in zebrafish, the animals develop smaller heads, which is one of the major symptoms of the human disease. Image shows a control zebrafish larva (top) and one in which expression of rpl10 was suppressed (bottom), resulting in normal body length but a proportionately smaller head. From Brooks and Wall et al.
———
Citations:
Brooks S.S., C. Golzio, D. W. Reid, A. Kondyles, J. R. Willer, C. Botti, C. V. Nicchitta, N. Katsanis & E. E. Davis (2014). A Novel Ribosomopathy Caused by Dysfunction of RPL10 Disrupts Neurodevelopment and Causes X-Linked Microcephaly in Humans, Genetics, 198 (2) 723-733. DOI: http://dx.doi.org/10.1534/genetics.114.168211 http://www.genetics.org/content/198/2/723.full
Hieter P. (2014). Understanding Rare Disease Pathogenesis: A Grand Challenge for Model Organisms, Genetics, 198 (2) 443-445. DOI: http://dx.doi.org/10.1534/genetics.114.170217
http://www.genetics.org/content/198/2/443.full
Johnston M. (2014). Humans as a Model Organism: The Time Is Now, Genetics, 198 (2) 441-441. DOI: http://dx.doi.org/10.1534/genetics.114.170225
http://www.genetics.org/content/198/2/441.full































