Check out the June issue of GENETICS by looking at the highlights or the full table of contents!
This Month’s Centennial Articles
Joshua Lederberg on bacterial recombination, pp. 613–614
Mark Johnston
GENETICS Editor-in-Chief Mark Johnston introduces Joshua Lederberg’s GENETICS Classic Gene recombination and linked segregations in Escherichia coli, which describes the first genetic analysis of bacteria. Lederberg’s discovery that E. coli is sexual allowed him to transform bacteria from a biological curiosity into an indispensible tool for modern genetics.
Curt Stern on somatic crossing over, pp. 615–616
James A. Birchler
GENETICS and G3 Associate Editor James A. Birchler introduces Curt Stern’s GENETICS Classic Somatic crossing over and segregation in Drosophila melanogaster. In this 106-page stemwinder, Stern describes his accidental discovery of the first example of somatic (mitotic) crossing over.
Massively parallel genetics, pp. 617–619
Jay Shendure and Stanley Fields
We can now conduct genetic analyses at a scale scarcely imaginable a decade ago. But the ease of genotyping has exposed the limits of genetic analyses, particularly as they have been applied to human phenotypes. GENETICS Associate Editor Jay Shendure and GSA President and GENETICS Associate Editor Stanley Fields highlight some of these limitations and argue that massively parallel approaches to experimentally measure the functional consequences of individual variants will advance our ability to interpret human genomes.
A brief history of Schizosaccharomyces pombe research: a perspective over the past 70 years, pp. 621–629
Peter A. Fantes and Charles S. Hoffman
Since its humble start as a model organism in two European laboratories in the 1940s and 1950s, the fission yeast Schizosaccharomyces pombe has grown to become one of the best-studied eukaryotes today. Peter A. Fantes and Charles S. Hoffman outline the way interest in S. pombe developed and spread from Europe to Japan, North America and elsewhere from its beginnings up to the first International Meeting devoted to this yeast in 1999.
—————————————————–
Functional divergence of the nuclear receptor NR2C1 as a modulator of pluripotentiality during hominid evolution, pp. 905–922
Jennifer L. Baker, Katherine A. Dunn, Joseph Mingrone, Bernard A. Wood, Beverly A. Karpinski, Chet C. Sherwood, Derek E. Wildman, Thomas M. Maynard, and Joseph P. Bielawski
To investigate the role of nuclear receptors (NRs) in hominid evolution, Baker et al. analyzed selection on all 48 human NRs. They identified NR2C1 as a candidate for contributing to the development of large primate brains. To evaluate this hypothesis, they inferred the sequence of NR2C1 for the ancestor of human and chimpanzees, and synthesized and expressed the ancestral protein in the laboratory. Functional comparison of the ancestral and modern proteins indicate they differ in several aspects of stem cell regulation affecting neuronal differentiation. The authors propose that NR2C1 could be associated with characteristics that distinguish humans from the other great apes.
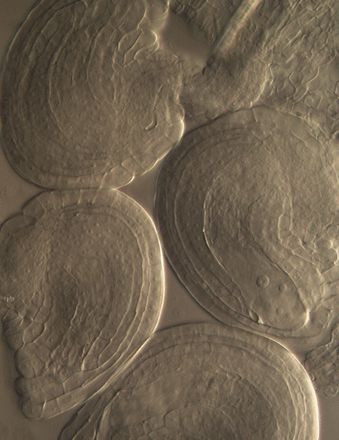
Cross-talk between sporophyte and gametophyte generations is promoted by CHD3 chromatin remodelers in Arabidopsis thaliana, pp. 817–829
Benjamin Carter, James T. Henderson, Elisabeth Svedin, Martijn Fiers, Kyle McCarthy, Amanda Smith, Changhua Guo, Brett Bishop, Heng Zhang, Tjitske Riksen, Allison Shockley, Brian P. Dilkes, Kim Boutilier, and Joe Ogas
The developmental complexity of multicellular organisms can obscure whether phenotypes resulting from loss of a ubiquitous regulator reflect the role of that factor in every tissue or only one. Carter et al. demonstrate that multiple reproductive defects associated with loss of a CHD chromatin remodeler in Arabidopsis result from loss in a distinct compartment, the maternal sporophyte. They also reveal opposing roles for two CHD remodelers in determining seed size, which acts as a novel trans-generational determinant of developmental identity in seedlings. These analyses illustrate the importance of considering non-autonomous mechanisms when investigating the contribution of an epigenetic regulator to a complex developmental process.
Evolution of schooling behavior in threespine sticklebacks is shaped by the Eda gene, pp. 677–681
Anna K. Greenwood, Margaret G. Mills, Abigail R. Wark, Sophie L. Archambeault, and Catherine L. Peichel
Greenwood et al. used transgenic approaches to show that natural variation in schooling behavior between two populations of threespine sticklebacks is caused by the gene Ectodysplasin. This work provides a rare example of a genetic change that underlies the evolution of vertebrate behavior in the wild.
The genetic cost of Neanderthal introgression, pp. 881–891
Kelley Harris and Rasmus Nielsen
Previous studies have shown that Neanderthals were inbred compared to humans, and also that Neanderthals interbred with the ancestors of people living outside Africa today, contributing 1 to 4% of their modern gene pool. We use simulations to show that Neanderthal inbreeding likely caused accumulation of harmful mutations, and they may have had 40% lower fitness than humans. Many of these harmful mutations are predicted to persist in modern human populations, although most have been eliminated by natural selection in humans since the time of interbreeding.
Unexpected roles for ciliary kinesins and intraflagellar transport proteins, pp. 771–785
Niedharsan Pooranachandran and Jarema J. Malicki
Cilia are essential for the function of sensory organs throughout the animal kingdom. Cilia formation and function depends on kinesin 2 family motors and on intraflagellar transport (IFT) proteins, which mediate transport inside the ciliary shaft. The authors show that both the main ciliary motor (heterotrimetic kinesin 2) and IFT proteins display unexpected functions in zebrafish photoreceptors, suggesting a much broader role than previously assumed. These functions are essential for photoreceptor cilia formation and photoreceptor survival. Surprisingly, however, sensory cristae in the ear do not require this motor for some aspects of ciliogenesis and the homodimeric kinesin 2 is not necessary for cilia morphogenesis in all tissues examined thus far.
Efficient genome-wide sequencing and low-coverage pedigree analysis from noninvasively collected samples, pp. 699–714
Noah Snyder-Mackler, William H. Majoros, Michael L. Yuan, Amanda O. Shaver, Jacob B. Gordon, Gisela H. Kopp, Stephen A. Schlebusch, Jeffrey D. Wall, Susan C. Alberts, Sayan Mukherjee, Xiang Zhou, and Jenny Tung
Methods for genotyping non-invasively collected samples have remained largely unchanged for the past 20 years. Snyder-Mackler et al. developed an efficient protocol for genome-wide capture of endogenous DNA from non-invasively collected samples and coupled it with a computational approach to reconstruct pedigree links. The enrichment method increased endogenous DNA up to 40-fold, and the computational approach reconstructed near-perfect pedigree relationships even with extremely low-coverage sequencing. These methods will enable research on natural populations to again take a major leap forward, into the genomic era.
Transcript isoform variation associated with cytosine modification in human lymphoblastoid cell lines, pp. 985–995
Xu Zhang and Wei Zhang
Natural variation of DNA methylation and its correlation with gene expression variation has been extensively studied. The extent of correlation between DNA methylation and alternative transcript isoforms among individuals and the underlying mechanisms, however, remain unclear. This study explored these aspects using genome-wide data derived from human lymphoblastoid cell lines from unrelated individuals. The study demonstrated a prominent effect of cytosine modification variation on transcript isoform spectrum over gross transcript abundance, and revealed epigenetic contributions to diseases that were mediated through cytosine modification-specific TIV.
Accurate profiling of gene expression and alternative polyadenylation with whole transcriptome termini site sequencing (WTTS-Seq), pp. 683–697
Xiang Zhou, Rui Li, Jennifer J. Michal, Xiao-Lin Wu, Zhongzhen Liu, Hui Zhao, Yin Xia, Weiwei Du, Mark R. Wildung, Derek J. Pouchnik, Richard M. Harland, and Zhihua Jiang
Construction of next-generation sequencing (NGS) libraries for transcriptome analysis involves RNA manipulation, which contributes to noisy, biased, and artifactual data. To minimize such problems, Zhou et al. developed a whole transcriptome termini site sequencing (WTTS-seq) method which enriches both polyA+ RNA and cDNA, adds 5′ and 3′ adaptors in one step, pursues strand sequencing, and profiles both gene expression and alternative polyadenylation (APA). The technique can also detect shorter transcripts and reduce false discovery of differentially expressed genes.