Multiparental populations (MPPs) have brought a new era in mapping complex traits, as well as new analytical challenges. To face these challenges and encourage innovation, the GSA journals launched the ongoing Multiparental Populations series in 2014. This month’s issues of GENETICS and G3 feature a bumper 16 MPP articles, timed to celebrate a new easy-to-use site for browsing the series. In line with our goal of encouraging communication across disciplinary boundaries, the “MPP People” profiles aim to introduce series authors working in a wide range of systems.
When MD/PhD student Andrew Morgan left Fernando Pardo-Manuel de Villena’s lab for clinical training earlier this year, his mentor took the loss philosophically.
“It’s actually nice to discover how much you depend on a really great student,” says Pardo-Manuel de Villena. “When they leave, there’s a big hole in the lab, and you feel that something is not working properly. But that’s a good sign of someone who is really exceptional.”
Morgan’s PhD work has spanned the diversity of Pardo-Manuel de Villena’s research program. At its heart is the study of evolutionary biology using experimental mouse genetics. “We’re trying to understand them as mice—not just some animal we use in the lab—and we’re learning how genetic variation can inform experimental design with lab strains,” says Morgan.

Andrew P. Morgan, University of North Carolina
During his undergraduate training in statistics, Morgan became interested in genetics, in part because of its elegant basic principles. Hoping to find a career where he could both solve problems and help people, he enrolled in the MD/PhD program at the University of North Carolina.
In his PhD rotations, Morgan was drawn to Pardo-Manuel de Villena’s group by the combination of bioinformatics, evolutionary genetics, and experimental genetics—along with the lab head’s boundless intellectual energy and supportive mentorship. Morgan flourished in this environment.
“He started as a fairly well-rounded grad student, but he went from there to writing his own papers—he has a paper in G3 where he’s the only author—to taking responsibility for starting and completing new projects and new collaborations to becoming a go-to person for many people in the department,” says Pardo-Manuel de Villena.
Much of his work has involved two powerful mouse resources, the Collaborative Cross (CC) and Diversity Outbred (DO) populations, which were designed to bring wild genetic variation into the the controlled environment of the lab.
The CC population was founded by crossing eight parental strains that incorporated diversity from both lab and wild-derived mice across three sub-species. The offspring were then inbred to generate many different stable inbred lines with approximately balanced genetic contributions from all of the founders. Each of these lines is a random genetic mosaic of the founders, but individuals within a line share a largely identical genome of a known genotype. The result is a unique resource for mapping QTLs and performing other genetic experiments in mice.
The DO population is a complementary resource created by randomly crossing 300 individuals from an early stage of the CC project. Instead of a fixed number of inbred lines with reproducible genomes, each DO mouse has a distinct and unique genome. This has yielded an outbred population with equivalent genetic diversity and allele frequencies to the CC, but with the founder genotypes more extensively recombined, allowing genetic mapping to a higher resolution than possible with standard QTL mapping or with the CC population.
In a paper published this month in GENETICS in the MPP series, Morgan and colleagues used genotype data from almost 7,000 DO mice to construct a very dense recombination map to study the influence of genetic variation on meiotic recombination. This huge pool of genotypes came from data shared by many research groups who have used the DO population in their experiments.
In mammals, meiotic crossovers are concentrated in small regions of the genome known as hotspots. The new DO analysis, covering more than 2.2 million crossover events, revealed that recombination is strongly suppressed in larger, spatially clustered coldspots that together make up about 12% of the observable genome. To investigate further, the team analyzed the whole-genome sequences of more than 200 DO mice and demonstrated that the recombination coldspots are strongly associated with segregating copy number variants. This work has established the importance of common structural variants in shaping recombination patterns and the utility of multiparental populations for bridging the gap between fine-scale and large-scale genetic mapping.
As well as performing mapping studies using the CC and DO mice, Morgan has played key roles in the development of new tools for their use, including mouse genotyping arrays optimized for subspecies-level genetic diversity and other analytical and bioinformatic resources.
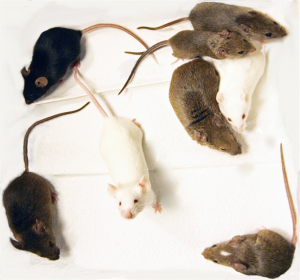
The eight founder strains of the Collaborative Cross mouse population. Photo: Brynn Voy, Oak Ridge National Laboratory.
During the epic cross-lab, cross-continental generation of the CC inbred lines, a number of unexpected observations have proven important. One was the unearthing of genetic incompatibilities between subspecies that greatly complicated the logistics of strain development but also revealed new insights into male fertility in mammals (see this month’s “On the Cover” post). A second curious result was that the allele frequency of a region on chromosome 2 was steadily drifting upward over the generations. Another grad student in the Pardo-Manuel de Villena lab, John Didion, showed that this evolution was due to a phenomenon known as meiotic drive. This means an allele was being preferentially transmitted during meiosis, violating Mendel’s law of random segregation.
Working together, Didion and Morgan found that the allele was a copy number expansion prevalent in wild populations of one of the subspecies used to found the CC. Not only had the allele repeatedly swept through lab populations like the CC, the same thing appeared to have happened in wild populations, all in spite of the fact that inheriting the allele seems to hamper reproductive success. Female mice heterozygous for this allele tend to have smaller litters in the lab than either homozygote–likely due to some degree of embryonic lethality associated with meiotic drive. This is a fascinating example of a selfish element that succeeds entirely through chromosomal mechanics, despite harming the fitness of individuals that carry it. “It bends several of the rules of how genetics is supposed to work,” says Morgan.
The discovery is not only relevant to evolutionary biologists: identifying this genetic hitchhiker allowed it to be purged from the DO population, restoring Mendelian transmission ratios to chromosome 2, where the gene resides, and preventing any potential analytical artefacts in future DO experiments.
After a productive PhD experience, Morgan is starting again at the bottom in his clinical training. “It’s kind of like going back to zero,” he says. But Pardo-Manuel de Villena is confident he will do as well in the clinic as he did in his research training. “He is smart, he has initiative, he is mature, and he is kind. That is a great combination.”
Read other MPP People profiles.
Browse the GSA Journals MPP series.
MPP AUTHOR:
Andrew P. Morgan, University of North Carolina
MPP ARTICLES:
Structural Variation Shapes the Landscape of Recombination in Mouse
Andrew P. Morgan, Daniel M. Gatti, Maya L. Najarian, Thomas M. Keane, Raymond J. Galante, Allan I. Pack, Richard Mott, Gary A. Churchill, and Fernando Pardo-Manuel de Villena
Genomes of the Mouse Collaborative Cross
Anuj Srivastava, Andrew P. Morgan, Maya L. Najarian, Vishal Kumar Sarsani, J. Sebastian Sigmon, John R. Shorter, Anwica Kashfeen, Rachel C. McMullan, Lucy H. Williams, Paola Giusti-Rodríguez, Martin T. Ferris, Patrick Sullivan, Pablo Hock, Darla R. Miller, Timothy A. Bell, Leonard McMillan, Gary A. Churchill, and Fernando Pardo-Manuel de Villena
John R. Shorter, Fanny Odet, David L. Aylor, Wenqi Pan, Chia-Yu Kao, Chen-Ping Fu, Andrew P. Morgan, Seth Greenstein, Timothy A. Bell, Alicia M. Stevans, Ryan W. Feathers, Sunny Patel, Sarah E. Cates, Ginger D. Shaw, Darla R. Miller, Elissa J. Chesler, Leonard McMillian, Deborah A. O’Brien, and Fernando Pardo-Manuel de Villena
Lisa E. Gralinski, Vineet D. Menachery, Andrew P. Morgan, Allison L. Totura, Anne Beall, Jacob Kocher, Jessica Plante, D. Corinne Harrison-Shostak, Alexandra Schäfer, Fernando Pardo-Manuel de Villena, Martin T. Ferris, Ralph S. Baric
Diversity Outbred Mice at 21: Maintaining Allelic Variation in the Face of Selection
, , , , , , , , , , , , and
The Evolutionary Fates of a Large Segmental Duplication in Mouse
, , , , , , , , , , and
The Mouse Universal Genotyping Array: From Substrains to Subspecies
, , , , , , , , , , and
argyle: An R Package for Analysis of Illumina Genotyping Arrays
Andrew P. Morgan
, , , , , and
, , , , and































