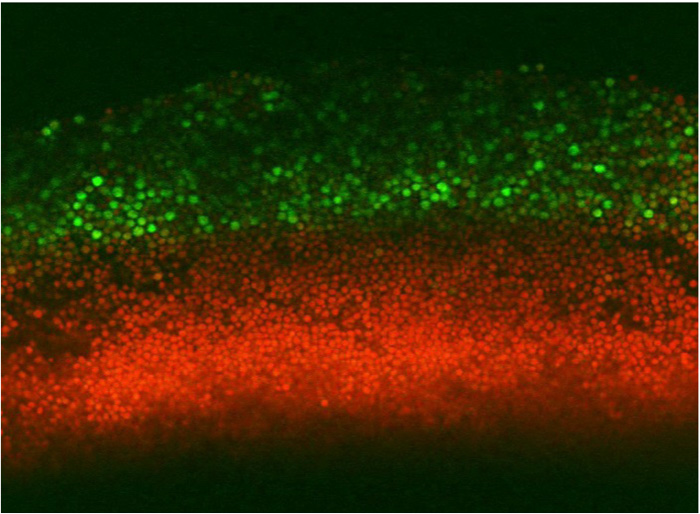
Compared to a human, a tree, or a jellyfish, the single-celled yeast might seem like a loner. Multicellular organisms like plants and animals are complex co-operative structures made of many specialized cell types, while a single yeast cell can survive and proliferate without the help of others. But although you might think of yeast as growing in the featureless soup of a lab culture, wild yeast typically live a more structured life—in colonies.
These colonies are dense communities of cells growing on a solid surface, and in recent years it has become clear that —like multicellular organisms— these colonies are composed of different cell types organized into patterns. But unlike for plants and animals, it remains unclear how this organization arises, or why it exists. New research in the latest issue of GENETICS reports on an apparent division of labor between two cell layers in yeast colonies, and suggests how this striking pattern may give the colony the flexibility to reproduce in diverse environments.
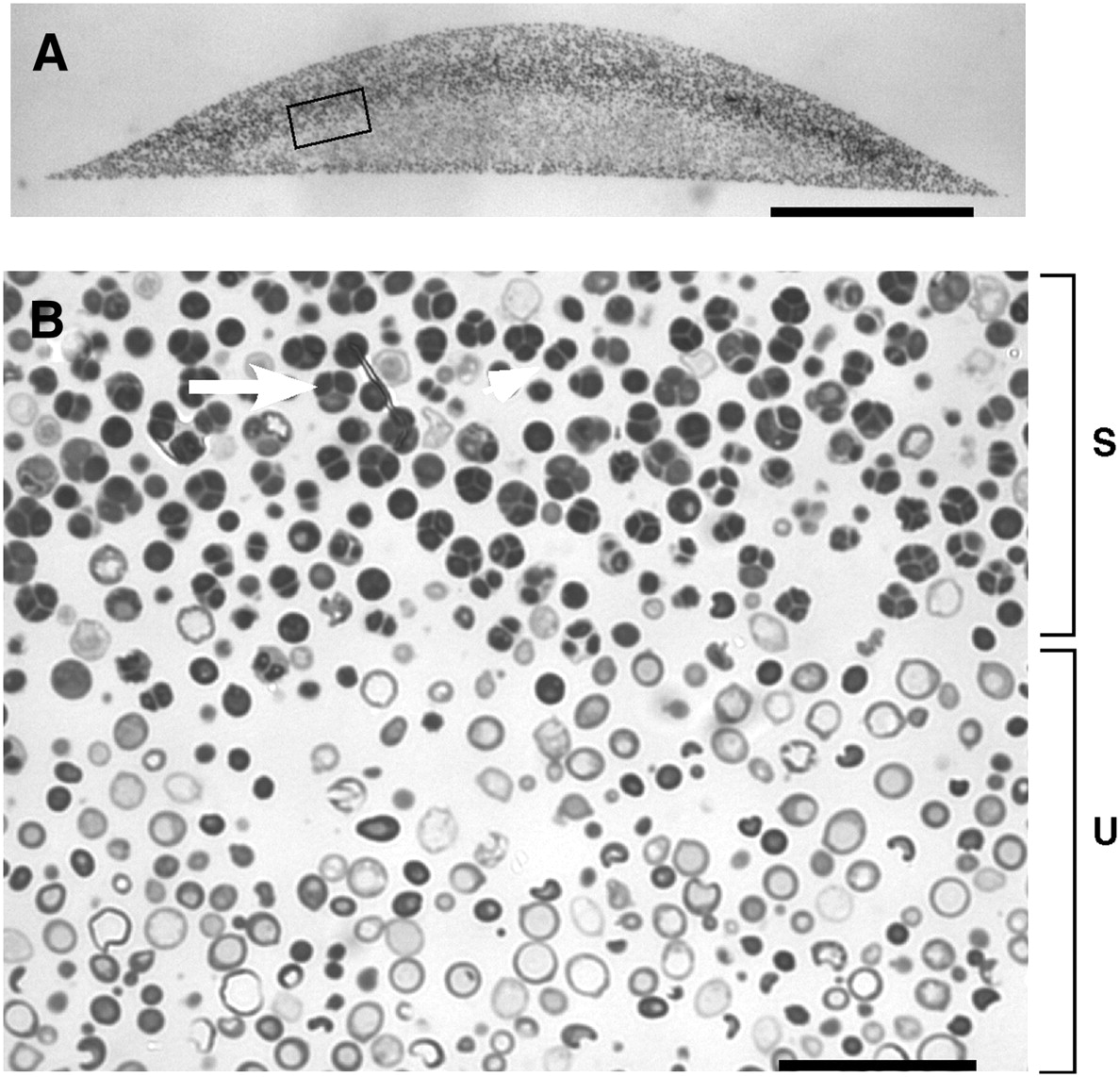
Spores in yeast colonies. (A) Cross section of embedded colony. (B) Higher magnification. The region of sporulation is indicated by a bracket marked “S”, and the unsporulated region indicated by a bracket marked “U”. The arrow indicates a tetrad asci (four spores) and the arrowhead indicates a dyad asci (two spores). From Piccirillo et al. 2010
The pattern investigated by Piccirillo et al. was the distribution of spores within colonies. Haploid spores are formed from diploid cells by meiosis, the first step of sexual reproduction. In a diploid colony, meiosis and sporulation begin in a sharp horizontal band of cells cutting through the center of the colony. In previous work, this layer was found to expand up to the top of the colony over several days, driven by progressive pH changes of the medium and activation of an alkali response signaling pathway. A second layer of spores also forms right at the bottom of the colony, but doesn’t expand in this way. This arrangement is not just a laboratory curiosity; the same distinctive organization is observed in yeast species isolated from the wild. This suggests the pattern may have been shaped by natural selection, contributing to reproductive success of the colony.
But if this particular pattern has an adaptive function, what could it be?
The authors started out by looking for genes important for sporulation only in colonies. They screened the yeast mutant collection for strains that sporulated poorly when grown as a colony, but which were able to sporulate to normal levels in liquid cultures (where spatial patterning is not possible because the yeast are in constant motion). They found that colony sporulation depends on the MAP kinase Cell Wall Integrity (CWI) pathway, which regulates remodeling and growth of the tough yeast cell wall.
Further investigation showed that the CWI pathway played a crucial, but indirect role in triggering sporulation in the top layer of spores. One to two days before this layer was established, a CWI pathway transcription factor, Rlm1, was activated in a separate band of cells underneath the cells that would later sporulate. Rlm1 activity in these underlying cells stimulated transcription of IME1, the master regulator that initiates meiosis, but only in the cells above them. The Rlm1-expressing cells also became more permeable to large molecules during development. This change in permeability may reflect cell wall remodeling that releases compounds stimulating sporulation in other cells.
Piccirillo et al. propose that the RLM1-expressing cells serve as a “feeder” layer, supplying nutrients to the sporulating cells above. This may be necessary because sporulation only occurs under starvation conditions, yet consumes considerable energy. Setting aside a portion of cells as feeders could allow the colony to optimize the distribution of scarce resources by adjusting the proportion of feeding and sporulating cells according to environmental conditions. Consistent with this idea, when the colonies were grown under suboptimal conditions of low carbon availability or high temperature, the feeder layer was proportionally bigger. Crucially, sporulation rates under these conditions were not only lower, sporulation was even more dependent on Rlm1 (and presumably feeder cells) than usual.
The authors refer to their hypothesis as differential partitioning-environmental buffering, and suggest that partitioning cells into different fates according to prevailing conditions could be a strategy used by many other types of cell coalitions — whether microbial communities, humans, trees, or jellyfish.
CITATIONS
Piccirillo, S., Morales, R., White, M., Smith, K., Kapros, T., & Honigberg, S. M. (2015). Cell Differentiation and Spatial Organization in Yeast Colonies: Role of Cell-Wall Integrity Pathway. Genetics, 201(4), 1427-1438 doi:10.1534/genetics.115.180919
http://www.genetics.org/content/201/4/1427
Piccirillo, S., White, M. G., Murphy, J. C., Law, D. J., & Honigberg, S. M. (2010). The Rim101p/PacC pathway and alkaline pH regulate pattern formation in yeast colonies. Genetics, 184(3), 707-716. doi: 10.1534/genetics.109.113480