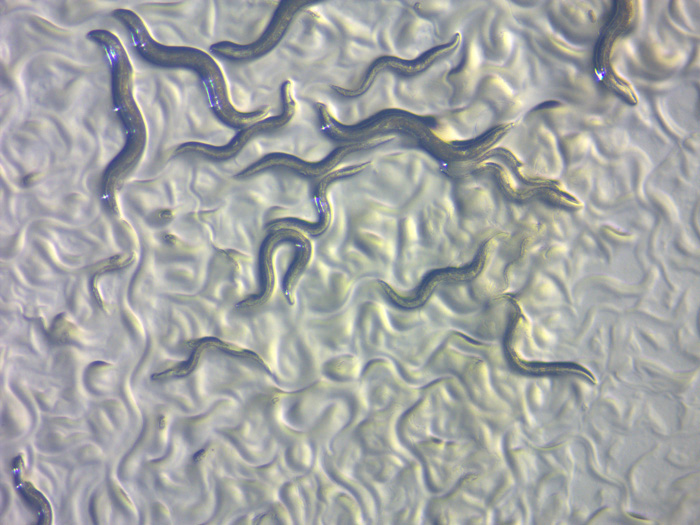
These worms are as long as a pencil’s tip and only just visible without a microscope. They are among the smallest multicellular animals, but they still have complex organ systems. They are Caenorhabditis elegans, one of the most important organisms in modern biology and a key to understanding the most basic molecular processes of life. Work on these tiny roundworms is helping geneticists unravel how all animals morph from single-celled embryos into fully-grown adults.
This tiny worm’s journey to prominence was kickstarted by a landmark paper published by Sydney Brenner in the May 1974 issue of GENETICS. As part of the GENETICS’ Centennial celebrations, Bob Goldstein introduced this paper as a GENETICS Classic. He reflects on how Brenner chose C. elegans for the study of complex phenomena like behavior and development. Brenner was looking for a new model organism at an exciting time for molecular genetics; the structure of DNA, the existence of mRNA, and the genetic code had all just been uncovered. Building upon this backbone of fundamental discoveries, the field was just starting to tackle more complex biological processes. A new approach was needed.
Brenner wasn’t the first scientist to think of C. elegans. The cultures that he began working with in the 1960s were originally isolated from an English compost pile. C. elegans is a species of roundworm. Though they are so small the average person doesn’t notice them, roundworms are thought to be the most abundant animal group, making up 80% of all individual animals living on land and 90% living in water. Roundworms are found in nearly every habitat imaginable, including the depths of the ocean and the bottom of a kilometers-deep gold mine. Many cause human diseases like hookworm. But C. elegans is harmless; it lives in soil and rotting fruits and vegetables where it feeds on bacteria.
Part of the power of C. elegans as a model system is how easy it is to raise them in the lab. They are very small and can easily be grown on petri dishes in large numbers, and they have a generation time of only a couple of days. For experiments that require examining multiple generations, this speeds up the process immensely. Their hardiness is also a benefit—living cultures of C. elegans can be frozen and then thawed later for further study. The mating system of C. elegans also makes them particularly well suited for genetics. They exist primarily as self-fertilizing hermaphrodites, so maintaining homozygous lines is very easy. Males do occur rarely, and they can be used to perform crosses to generate new hermaphrodite lineages.
Tiny C. elegans is also transparent, so its anatomy is easy to observe with a relatively low power microscope. This simplicity is a large part of what drew Brenner to use C. elegans for his work on complex phenotypes. It is a multicellular eukaryote, but one that is small and simple enough to have its genetics and physiology thoroughly dissected and understood in the lab. In the introduction to his 1974 paper “The genetics of Caenorhabditis elegans,” Brenner outlines how his ultimate goal is to understand the complete genetics and structure of the nervous system. C. elegans is perfectly suited for this; it has fewer than a thousand cells, half of which are neurons. In fruit flies, which have billions of neurons, this undertaking would be impossible. Brenner then describes the first steps toward his goal—a massive mutational screen designed to identify as many genes as possible.
C. elegans eventually proved to be the powerful model Brenner had hoped for. By 1977, the developmental fate of every postembryonic cell was known. One groundbreaking consequence of this work was the identification of the genes involved in programmed cell death. The 2002 Nobel Prize in Medicine was given to Sydney Brenner, H. Robert Horvitz, and John E. Sulston for their use of C. elegans to illuminate the genetic regulation of development and programmed cell death. Many of the pathways that regulate these processes in C. elegans are found all across the tree of life.
In 1986, Brenner achieved his goal of describing the connectivity and structure of all 302 neurons in C. elegans. This remains the only fully described “connectome” of any species, though the scientists of the Human Connectome Project are working hard to achieve the same for the more complicated human brain. C. elegans has continued to lead the way in many other research areas. In 1998, C. elegans was the first multicellular species to have its genome sequenced. The use of RNA interference to silence gene expression was also pioneered in C. elegans. This allowed the creation of large-scale screens to decrease expression of each gene in the genome one by one and then observe the resulting phenotypic changes—exactly the kind of exhaustingly thorough genetic study that embodies Sydney Brenner’s pioneering vision.
CITATIONS
Goldstein, B. (2016). Sydney Brenner on the Genetics of Caenorhabditis elegans. GENETICS, 204(1), 1-2. DOI: 10.1534/genetics.116.194084
http://www.genetics.org/content/204/1/1
Brenner, S. (1974). The genetics of Caenorhabditis elegans. GENETICS, 77(1), 71-94.