For every rule, there are exceptions. The same is true for how organisms organize their genomes. From how DNA is packaged in the nucleus to what genetic code is used for translating proteins, notable exceptions are found all across the tree of life. Most organisms use the standard genetic code. A few, such as ciliates, use a different code. Most organisms condense their DNA using histone proteins, packing strands of DNA into structural units called nucleosomes. A few don’t, the most notable exception being dinoflagellates, which, instead of using histones, supercoil their DNA into liquid-crystal condensates. The typical mammalian genome is between 1.6–6.3 billion nucleotides in length. For other types of organisms, such as the carnivorous plant genus Genlisea, their genomes are ultra-short with variable lengths due to a loss in non-coding regions.
These exceptions teach us what the rules are. With the advent of mass genetic sequencing capabilities and major sequencing initiatives such as the Earth Biogenome Project, we are learning more about the different types of genomic organization. Many of these defy our expectations, offering new insights about the rules for genomic organization, as well the selective pressures affecting genomic evolution.
Dinoflagellates: small organisms, big genomes
Source: Flickr
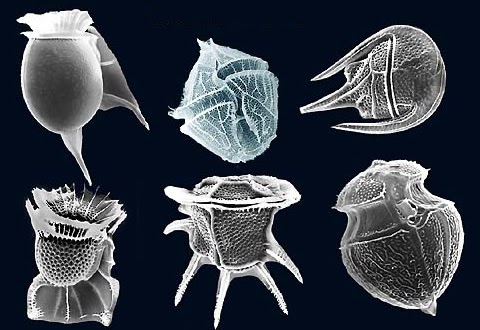
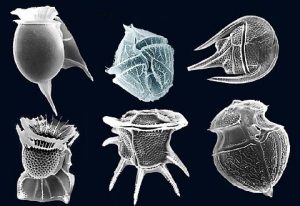
Dinoflagellates are marine algae which often make the news due to the phenomenon known as a “red tide.” A red tide occurs when dinoflagellates and other marine microorganisms reach high enough numbers to change the color of the water to a dark red color. It is toxic to humans, fish, mussels, and more. But despite this toxicity, dinoflagellates are also critical for the survival of any water environment and help with the growth of reef-building coral.
Dinoflagellates, which have genomes that are up to 80 times larger than the human genome, are an excellent organism for understanding the different ways of organizing DNA inside of the nucleus. Instead of nucleosomes, which are found in most eukaryotes, dinoflagellate DNA is supercoiled into liquid crystalline chromosomes (LCCs). These LCCs are permanently condensed DNA fixtures of the nucleus and even remain while dinoflagellate cells divide. Researchers are still working to uncover the mysteries of how these structures evolved, how they are created and maintained in the living dinoflagellate, and how LCCs impact the dinoflagellate life cycle.
Ciliates: mixing things up
Source: Robinson 2006
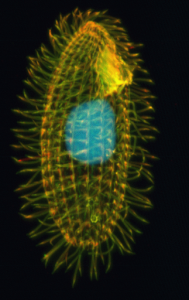
Ciliates are mostly unicellular protists that can be found in a variety of environments, ranging from marine and freshwater habitats to animal intestines. Their genomes defy many of the rules found in introductory textbooks. One rule many ciliate species don’t follow is using the “universal” genetic code. The four letters of DNA—A, T, C, and G—code for amino acids that build proteins in triplets called codons. The codon sequence ATG, for example, codes for the amino acid methionine. But three codons typically code for stop sequences that trigger the end of protein synthesis, such as the sequence TAA. In nearly all other living things, this sequence (TAA) is a stop signal, but in many ciliates, like Paramecium bursaria, this sequence codes for the amino acid glutamine instead. A recent study even suggests that one ciliate species, Condylostoma magnum, has a genetic code where all three stop sequences have been re-coded!
Perhaps the most remarkable feature of ciliates though is they possess two different types of nuclei, a trait called nuclear dimorphism. These two nuclei are both functionally and structurally distinct. The larger macronucleus controls the day to day functioning of the cell while the smaller micronucleus is strictly used for sexual reproduction. The genome of the macronucleus is derived from that of the micronucleus but looks dramatically different. For example, the micronucleus of the model ciliate Tetrahymena thermophila contains five chromosomes, but its macronucleus contains approximately 225 chromosomes. This striking difference in structure is because the macronuclear genome goes through major rearrangements during its transition from a micronucleus; multiple processes occur, such as editing out about one-third of the genome and making numerous copies of other parts.
The genomic gymnastics of ciliates have led to several major discoveries about genome biology and two Nobel prizes. The ends of chromosomes in eukaryotes have protective caps called telomeres, which are known to play important roles in health—including cancer and aging. The enzyme telomerase, which constructs telomeres, was first discovered and characterized in Tetrahymena. Genomic rearrangement is also a common occurrence in cancer cells, so studying how extreme genome rearrangements occur in ciliates may provide insight in their role in cancer as well. By showing us that the genetic code and genome structure is highly malleable, ciliates can help us understand that the language of life is not set in stone.
Genlisea: a tiny but mighty genome

Source: Darwiniana
Carnivorous plants are well-known members of the plant kingdom because of their fascinating ability to capture insects and digest their bodies for nutrients. Genlisea—named the “corkscrew plant” because they possess heavily twisted, modified leaves that trap organisms in the soil— is one such carnivorous plant that captures insects in their leaves for survival. Members of the genus Genlisea possess another feature that is just as unique: an ultra-small genome that is highly variable in size. G. aurea and G. margaratae, with genome sizes of 63.6 million base pairs and 63.4 million base pairs, respectively, have the smallest genomes. G. uncinata’s genome, at 1700 million base pairs, is 27 times larger. These genome sizes are a stark contrast to the largest known plant genome, Paris japonica, which has 150,000 million base pairs.
In addition to the large variability in genome size, Genlisea genomes also come with a high variation of chromosome number, with more derived species bearing a smaller number. Such small genomes within Genlisea can be attributed to genome contraction during its evolution through gene loss, and reduction of lengths of introns (non-coding DNA) and intergenic regions (stretches of DNA sequences in between genes) over time.
Even more interesting is that while gene loss and gene changes occur within the nuclear genome, the Genlisea genus has also experienced loss of genes in the chloroplast genome. An example of this is the ndh gene (NAD(P)H-dehydrogenase complex). This complex is involved in photosynthesis and stress response. It has also been speculated to have been related to the transition to terrestrial habitats. Researchers compared the chloroplast genome of terrestrial Genlisea species with closely related aquatic Utricularia species and observed that, compared to Utricularia, Genlisea ndh genes are either deleted, fragmented, or pseudogenized (DNA fragments that resemble functional genes but are nonfunctional). This finding of gene deletion is particularly insightful, because it opens a window to understanding how the evolution of ndh genes could be involved in the transition from water to land.
Scientists speculate that the occurrence of such dramatic changes in the genome by way of contraction, gene loss, and substitution are largely factors that facilitated the evolution of Genlisea’s unique insect-trapping ability. G. aurea does not actually contain roots, but instead possesses modified leaves that traps organisms. The lack of roots may be a factor that permitted the movement from water to land. Combined with its evolution from aquatic species to land, Genlisea is a truly unique plant genus.
Useful insights from unexpected places
These are just a few examples of organisms with genomes that defy expectations. By studying them, scientists have learned about the different ways DNA can be organized and changed, which has generated major shifts in our understanding of life around us. The recent boom in genome sequencing technologies and major projects like the Earth BioGenome initiative will undoubtedly find even weirder and fascinating genomes, yielding even more insights.
Written by members of the Early Career Leadership Program:
Zach I. Grochau-Wright, PhD, Department of Biology, The College of New Jersey
Jessica M. Vélez, PhD , Bredesen Center for Interdisciplinary Studies, University of Tennessee, Knoxville
Adelita D. Mendoza, PhD, Department of Developmental Biology, Washington University